Researchers at the Johns Hopkins Kimmel Cancer Center are advancing a novel blood testing technology known as GEMINI (Genome-wide Mutational Incidence for Non-Invasive detection of cancer).
By merging genome-wide sequencing of single DNA molecules discarded by tumors with machine learning (ML), the test may enable earlier detection of lung and other cancers.
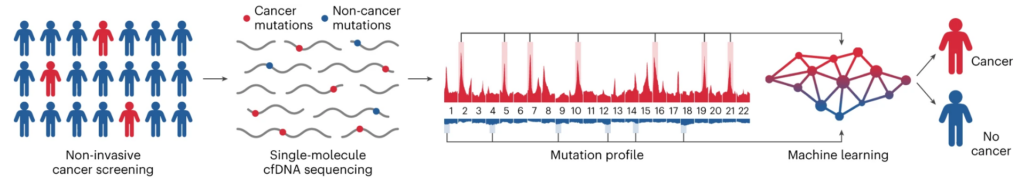
The GEMINI test analyzes cell-free DNA (cfDNA) fragments shed by tumors, using ML to detect differences in cancer and non-cancer mutation frequencies across the genome. Higher scores generated by the model suggest a higher probability of having cancer.
In lab tests, GEMINI was applied to computerized tomography images and detected over 90% of lung cancers, even at early stages. The study findings were published in Nature Genetics.
The researchers focused mainly on lung cancer detection in high-risk populations.
However, altered mutational profiles were also observed in cfDNA from patients with liver cancer, melanoma, and lymphoma, so this approach could be used across different types of cancers.
How the researchers created GEMINI
To develop GEMINI, the team studied genome sequences from 2,511 individuals across 25 different cancers.
Different tumor types featured different mutation profiles. The researchers found that genomic regions with a high mutation frequency were similar between tumor tissue and blood-derived cfDNA in patients with lung cancer, melanoma, or B cell non-Hodgkin lymphoma.
The GEMINI test was then applied to cfDNA from 365 individuals at high risk of lung cancer, producing higher scores in people with cancer than those without.
Researchers also examined integrating GEMINI with a previous test known as DELFI (DNA evaluation of fragments for early interception),
In total, GEMINI and DELFI correctly identified cancers 91% of the time in 89 samples.
Remarkably, the GEMINI test detected abnormalities in cfDNA mutation profiles years before standard diagnoses in seven patients with no detectable tumors at the time of blood collection.
Six tested positive using GEMINI and were later diagnosed with lung cancer from 231 to 1,868 days after samples were obtained.
Breakdown of the study
- Blood sampling: A blood sample is taken from a person who might be at risk for developing cancer. This sample contains cfDNA, tiny fragments of DNA that have been shed by cells in the body, including any potential tumor cells.
- DNA extraction and sequencing: The cfDNA is extracted from the blood sample and then sequenced. This means that scientists map out the order of the ‘building blocks’ that make up the DNA. This helps them identify any changes or mutations to the DNA.
- Analysis of DNA alterations: Each individual DNA molecule is analyzed for any sequence alterations, allowing the researchers to map out mutation profiles across the entire genome. In essence, they’re looking for patterns in the mutations that might suggest the presence of cancer.
- Machine learning: A machine learning model, which has been trained to recognize the difference between cancerous and non-cancerous mutation frequencies in different areas of the genome, is then used to analyze the mutation profiles. The machine learning model then gives a score ranging from 0 to 1, where a higher score indicates a higher likelihood of cancer.
- Further validation and testing: If the GEMINI score is high, suggesting the presence of cancer, additional tests such as computerized tomography imaging and the DELFI test (which detects changes in the size and distribution of cfDNA fragments across the genome) are used to confirm the diagnosis and detect the stage of cancer. This combination of tests has proven very effective, detecting over 90% of lung cancers in the study.
This is another intriguing study revealing the novel application of machine learning to medical diagnostics, this time analyzing cancers at the cellular level.
Larger clinical trials are required to validate the tool before it can become available for clinical use.
This week, an AI-supported breast cancer screening workflow vastly improved the speed and efficiency of assessing mammograms for cancer.